Skin Research Platform
Humanized 3D Skin models, Host-Skin Microbiome Interaction model
Skin Research Innovation

Complementary Technical Capabilities
We presents a comprehensive and state-of-the-art platform for research into human skin health. Our expertise spans the full R&D spectrum, including computational Safety & Sustainability by Design (SSbD), in vitro skin models for preclinical studies, high resolution molecular and multi-omics analyses, in vivo clinical research, and wearable technologies for continuous skin monitoring. These capabilities can be deployed individually or in combination.
Broad Application Areas
TNO enables partners from multinational companies to public institutions to accelerate skin focused innovation through tailored consultancy, investigative research, collaborative innovation, and orchestration of multi stakeholder initiatives.
Partnership with Public and Private Organizations
With our Skin Research Platform, we offers pharma and cosmetic partners an unique opportunity to drive breakthroughs in skin health through an integrated, science driven, and future ready approach. By combining cutting edge technologies with deep multidisciplinary expertise and independent validation, we help organizations de-risk development, strengthen claims, and bring safe, effective, and sustainable innovations to market faster. Together, we can shape the next generation of skin health solutions for individuals, industry, and society.
Cell Assays - 3D Human Skin models (Animal Material Free)
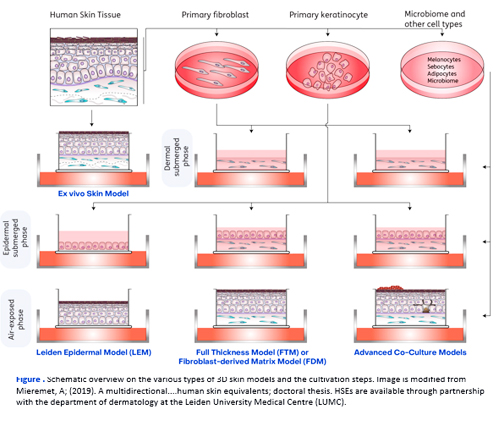
Human skin 3D models can be deployed to determine effects of intervention in a physiologically relevant setting. Various types of 3D models can be constructed, based on the level of throughput and complexity required in the research project.
Ex vivo Skin Model
The ex vivo skin model is based on cultivation of biopsies derived from elective plastic surgery. In the ex vivo skin model, the cellular organization and the extracellular matrix reflects that of in vivo human skin closely.
Human Skin Equivalents (HSEs) :
● Leiden Epidermal Model (LEM)
The LEM is a skin model based on cultivation of primary keratinocytes and reflect epidermal structure. During submerged phase, the keratinocytes proliferation and cover the insert fully, whereas during air-exposed phase differentiation occurs. After terminal differentiation, the LEMs represent all major epidermal differentiation layers. This model be upscaled for high-throughput screenings.
● Full Thickness Model (FTM)
The FTM entails a 3D skin models that consists of both fibroblasts and keratinocytes. The fibroblasts are cultivated in a collagen matrix, thereby mimicking the dermal compartment of the skin. In the FTM, fibroblast-keratinocyte interactions can be studied, as both cell types are present.
● Fibroblast-Derived Matrix Model (FDM)
In the FDM, also the two main cell types of the skin are cultivated. In the FDM, the fibroblast secreted and modulated the extracellular matrix, which thereby solely consists of cell-derived proteins. The FDM can be generated with specific fibroblasts subtypes that can reflect deeper dermal layers.
Advanced Co-Culture Models
Increased complexity is achieved with advanced co-culture models. Additional cell types can be introduced, which are melanocytes, sebocytes, or adipocytes. To determine host-microbiome interactions, bacteria, viruses, or fungi can be introduced topically. Next-generation advanced models include immunocompetent and skin-on-a-chip models.
Skin Microbiome
The skin microbiome is a diverse community of bacteria, fungi, viruses, and mites that live on the skin’s surface.
It plays a key role in protecting against harmful pathogens and supporting the skin’s immune function. When balanced, the skin microbiome helps maintain healthy skin, but disruptions can contribute to conditions like acne or eczema.
Strain collecting and culturing
Skin microbiome strain collection focuses on isolating representative microbes from healthy or diseased skin to enable studying isolated strains.
Culturing enables the expansion and phenotypic study of individual strains under controlled conditions. In-house whole-genome sequencing enables characterization of these strains at high resolution, revealing taxonomy, functional genes, and strain level variation.
Collected strains are stored in the TNO strain collection combined with host-related metadata and are accessible for R&D projects.
Skin microbiome characterization
Characterization of the skin microbiome as complex ecosystem is essential to understand its link with skin health. Several methods are developed for skin microbiome characterization.
Our host-depletion swab methods enrich microbial DNA by reducing human DNA contamination from skin samples. 16S rRNA gene sequencing profiles bacterial community composition and relative abundance at the genus or species level.
Shotgun metagenomics enables high resolution taxonomic assignment and direct assessment of microbial genes and pathways. Integrated bioinformatic data analysis reveals both microbial taxonomy and functional capacity across skin microbiome samples.
In vitro skin microbiome assay
An in vitro microbiome assay uses defined bacterial communities to model microbial interactions under controlled conditions.
It enables the study of bacterial growth, competition, and response to external factors such as nutrients or compounds.
Our bespoke assays provide reproducible insights into microbiome behavior while reducing the complexity of in vivo systems.
Strain engineering
Strain engineering of skin microbes involves modifying native skin bacteria to enhance beneficial functions such as antimicrobial production or barrier support.
These engineered strains are explored for therapeutic and cosmetic applications while prioritizing safety, stability, and host compatibility.
Host-Microbe Interaction Models
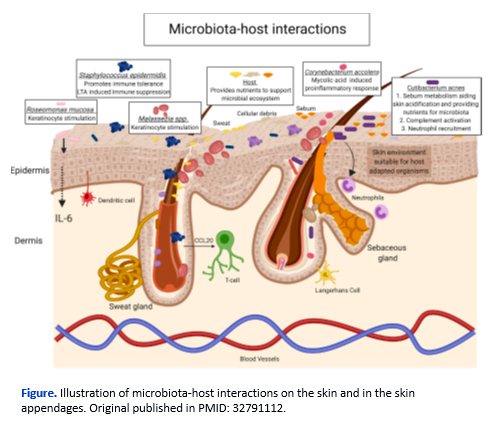
Determination of the interactions between the microbiome and the human host can be performing using co-culture models. At TNO, multiple types of host-microbe interaction models have been developed and applied in scientific research.
These include 2D models based on primary cells enriched with bacteria, fungi, and/or viruses. Examination of physiological relevant interactions on the skin of the microbiome can be performed with 3D models, which can be enriched with microbial mono-, or polycultures to model the microbiome.
Screenings in 2D cell-microbe assays
A skin host-microbe interaction model uses 2D cultures of keratinocytes, fibroblasts, or other skin cell types to represent the human host. These models are co-cultured with bacteria, fungi, or viruses to study direct microbial interactions with skin cells.
They enable controlled investigation of immune responses, barrier function, and microbial effects on skin physiology.
Validations with 3D host-microbe co-culture models
Three dimensional skin models, including reconstructed epidermis and full-thickness skin equivalents, more faithfully recapitulate tissue architecture and barrier function.
These systems incorporate multiple host cell types in a stratified context, enabling physiologically relevant host-microbe interactions. Colonization with bacteria, fungi, or viruses allows investigation of spatial organization and niche specific microbial behavior.
3D models capture complex immune and metabolic responses that are not accessible in 2D cultures.
As such, they provide a translational platform for studying skin biology, disease mechanisms, and microbiome-targeted interventions.
Tailored approaches for atopic dermatitis and acne
Modulation of 3D host microbe co-culture models enable disease specific interrogation of atopic dermatitis and acne by incorporating relevant skin architecture and barrier defects.
Selective colonization with pathobionts such as Staphylococcus aureus or phylotype-specific Cutibacterium acnes captures disease associated microbial dynamics.
These models facilitate mechanistic and translational studies of inflammation, host susceptibility, and microbiome targeted interventions.
Publication
Scientific Reports. March 2026
Advancing human skin models by integrating skin microbes for next-generation research.
Tissue Engineering Part A. Dec 2017
Recapitulation of Native Dermal Tissue in a Full-Thickness Human Skin Model Using Human Collagens.
Eur J Dermato. June 2017
Differential effect of extracellular matrix derived from papillary and reticular fibroblasts on epidermal development in vitro.
Mutagenesis. Mar 2011
Development and characterisation of an in vitro photomicronucleus test using ex vivo human skin tissue.
Toxicology in vitro. Aug 2008
Leiden reconstructed human epidermal model as a tool for the evaluation of the skin corrosion and irritation potential according to the ECVAM guidelines.

